 ,
,
1) Indication & Clinical Positioning
Primary indication:
- Rapid detection of MTBC DNA/RNA in specimens where rapid rule-in/rule-out can change isolation status or therapy.
- Pulmonary: sputum (spot/morning/induced), bronchoalveolar lavage (BAL), tracheal aspirates.
- Extrapulmonary: CSF, pleural fluid, lymph node aspirates/tissue, gastric aspirate, urine (GU TB), bone/soft tissue.
When it’s especially useful:
- Smear-negative or paucibacillary disease.
- Severe disease with high consequence of delay (e.g., TB meningitis).
- Prior antimicrobial exposure likely to suppress culture.
- Infection control decisions (airborne isolation, cohorting).
Relationship to other methods:
- Smear microscopy: fast but insensitive/specificity-limited.
- Culture: gold standard for viability and drug susceptibility; slower.
- Molecular NAAT: complements both fast etiologic evidence; may include resistance markers depending on assay.
2) Pre-Analytics: Specimens, Transport, and Biosafety
Collection & matrices
- Sputum/BAL: decontaminate/digest as per lab SOP (e.g., NALC-NaOH) before extraction for DNA assays.
- CSF: prioritize volume; concentrate if low biomass.
- Tissue/FNA: mince/homogenize in sterile buffer; avoid fixatives for molecular testing.
- Urine/pleural/ascitic/other fluids: centrifuge and process pellet.
Transport & storage
- Process promptly; keep cold if delays are expected.
- Avoid repeated freeze–thaw; use aliquots.
Biosafety
- Pre-inactivation steps in BSC-II with appropriate PPE.
- Physical separation of pre- and post-amplification areas.
3) Assay Design: Controls, Targets, and Thresholds
Targets
- MTBC-specific genomic regions (e.g., insertion sequences, multicopy regions, or species-discriminant loci).
- Optional modules for common resistance determinants (e.g., rifampicin hot-spot, isoniazid-associated loci) depending on the platform.
Controls (must-have)
- Extraction control (IC-extraction): monitors lysis, purification, inhibitors.
- Amplification control (IC-PCR): detects PCR failure independent of target.
- Positive control: low-copy MTBC template for run acceptance.
- Negative controls: non-template control (NTC) and negative extraction blank.
Interpretation thresholds
- Ct/quant cutoffs established during local verification/validation (matrix-specific).
- Decision tree should incorporate: Ct range, curve shape, replicate agreement, and control performance.
- For cartridge-based NAATs, use manufacturer’s result flags plus local QC acceptance criteria.
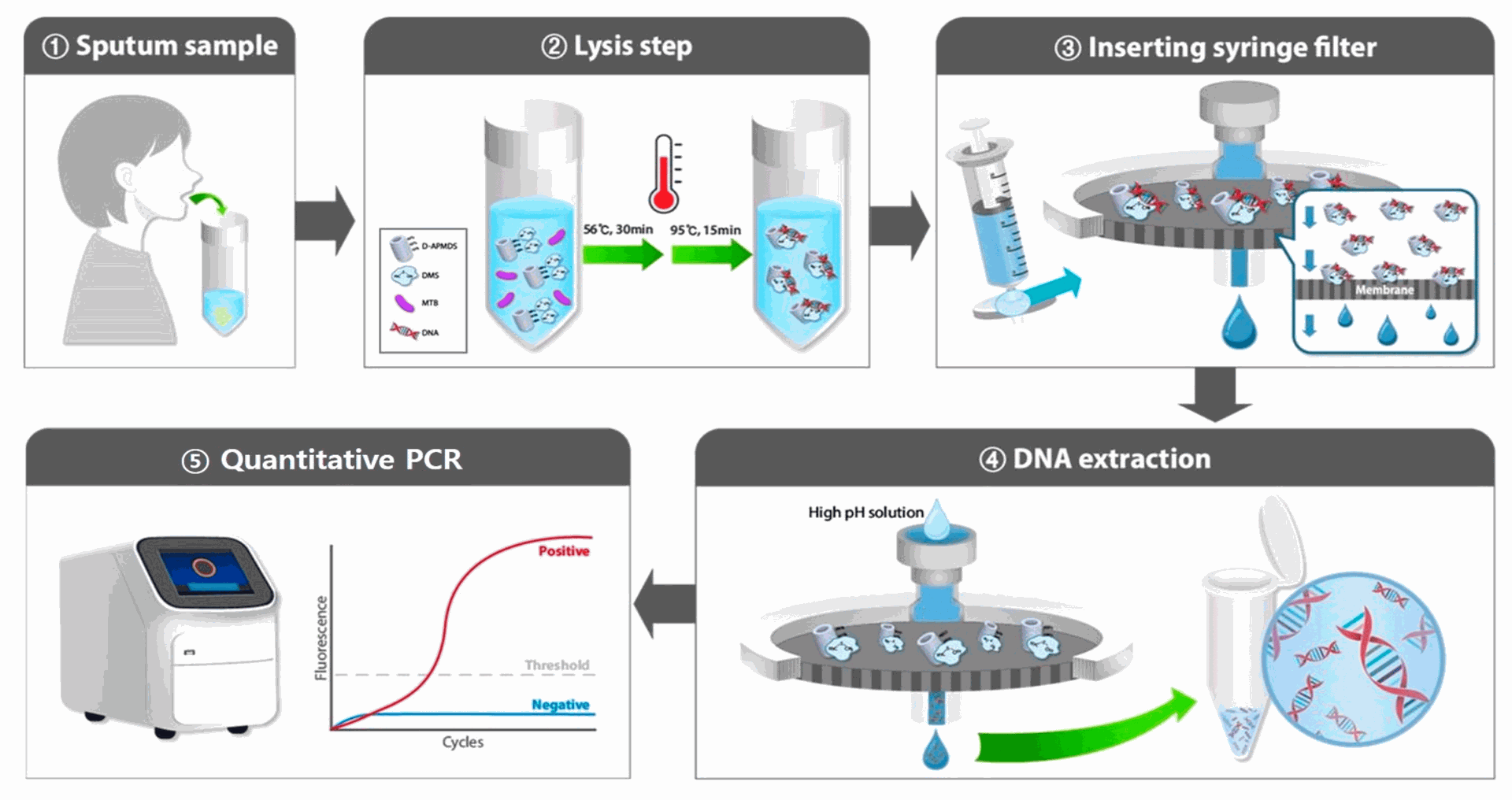
4) Workflow Overview
- Specimen processing (decontamination/digestion where applicable).
- Nucleic-acid extraction (silica membrane or magnetic beads).
- Amplification (real-time PCR, isothermal, or nested PCR depending on kit).
- Detection & call (Ct values/qualitative flags; optional melt/probe signatures).
- Result verification (control review, delta-Ct sanity checks, inhibition assessment).
- Report generation (qualitative call ± Ct/quant; adequacy and limitations).
Turnaround time: commonly same day for high-throughput labs; 2–24 h depending on batching and method.
5) Analytical Performance & Validation Plan
Key studies to complete locally (per matrix):
- LoD (limit of detection): serial dilutions around decision threshold.
- Inclusivity: panel of MTBC strains across lineages.
- Exclusivity: panel including NTM (non-tuberculous mycobacteria) and common respiratory flora.
- Precision/Reproducibility: inter-day, inter-operator, inter-instrument.
- Interference: blood, mucus, host DNA excess, medications (e.g., rifampicin carryover), and specimen preservatives.
- Carryover/Cross-contamination: high-positive next to negatives.
- Specimen stability: time/temperature studies.
Typical expectations (assay/platform-dependent):
Sensitivity: higher in smear-positive pulmonary disease; variable in extrapulmonary/low biomass.
Specificity: high with proper exclusivity and contamination controls.
Inhibition rate: track and manage via IC and re-extraction policies.
6) Result Interpretation Framework
Positive (MTBC detected):
- Meets Ct/curve acceptance and controls pass → Report detected.
- Flag if near-threshold; consider repeat on residual extract when clinically borderline.
- For extrapulmonary matrices, add a statement on lower biomass and need for clinical correlation.
Negative (MTBC not detected):
- Controls pass; no MTBC signal → Report not detected with sensitivity caveat (LoD, matrix).
- If inhibition (failed IC) → Invalid, recommend re-extraction or recollection.
Indeterminate/Repeat:
- Discordant replicates, atypical curves, or control anomalies → repeat testing; consider alternative matrix.
Resistance markers (if included):
- If the assay includes limited genotypic markers (e.g., rifampicin region), clearly state scope and that phenotypic DST or comprehensive genotyping may still be required.
7) Reporting: Clear, Actionable, and Audit-Ready
Minimum elements:
- Patient/specimen identifiers (per local policy), specimen type, collection date/time.
- Method summary (NAAT type, target class), assay/pipeline version.
- Qualitative result: “MTBC Detected / Not Detected / Invalid.”
- Optional Ct/quant metrics (assay-dependent) and IC status.
- Sample adequacy comment (e.g., IC recovered; no inhibition detected).
- Interpretation notes: clinical correlation required; culture/DST recommendations; isolation implications per local guideline.
- Limitations: LoD/matrix effects; does not assess viability; resistance scope if applicable.
Example snippet:
- Result: M. tuberculosis complex Detected
- Controls: IC-extraction PASS; IC-PCR PASS; NTC PASS
- Metrics (optional): Target Ct 33.1 (threshold ≤37 by validation)
- Adequacy: No inhibition detected; sample volume 1.5 mL post-concentration
- Interpretation: Molecular evidence of MTBC nucleic acid; correlate with clinical/radiographic findings. Send culture for DST. Maintain airborne precautions per policy.
- Limitations: Molecular detection does not confirm viability; resistance not assessed beyond the loci covered by this assay.
8) Quality System & Ongoing Monitoring
- EQA/Proficiency testing: enroll where available; track scores and CAPA.
- Run acceptance rules: control-based; lock and document.
- Contamination surveillance: trend NTC signals, environmental swabs if incidents arise.
- Change control: database/primer/probe updates, software versions, reagent lot changes.
- Training: competency assessment for specimen processing, inhibition troubleshooting, and report writing.
9) Common Pitfalls & Troubleshooting
- False negatives: low biomass, inhibitors, improper decontamination (over- or under-processing), delayed transport.
- False positives: amplicon carryover; mitigate with unidirectional workflow, dUTP/UNG where applicable, and strict area segregation.
- Borderline Ct: repeat on residual extract and consider clinical context; do not over-interpret single late-Ct without supportive evidence.