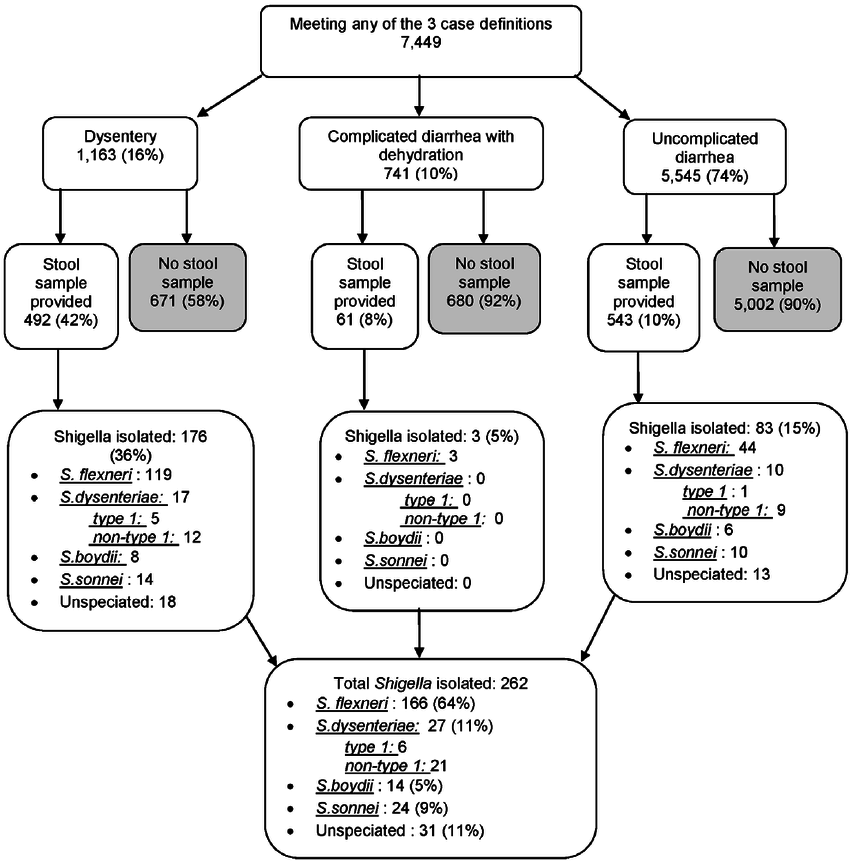
A 12-target diarrhea pathogen panel uses multiplex nucleic-acid testing (NAAT) to detect high-prevalence viral, bacterial, and protozoal enteric pathogens in a single run. The objective is fast rule-in/rule-out during suspected gastroenteritis, supporting infection control and clinical decisions especially in hospitals, long-term care, and outbreak investigations.
Coverage (Representative, non-exhaustive)
- Vir uses: Norovirus (GI/GII), Rotavirus A, Adenovirus (enteric types), Sapovirus, Astrovirus
- Bacteria: Salmonella enterica, Shigella spp./EIEC marker, Campylobacter spp., Clostridioides difficile toxin gene(s), Vibrio spp. (region-dependent), heat-labile/heat-stable toxin markers for ETEC/EAEC (as applicable)
- Protozoa: Giardia duodenalis, Cryptosporidium spp., Entamoeba histolytica
The exact 12 targets should reflect local epidemiology, travel patterns, and regulatory labeling.
vft
Gut Microbiota and Diarrhea
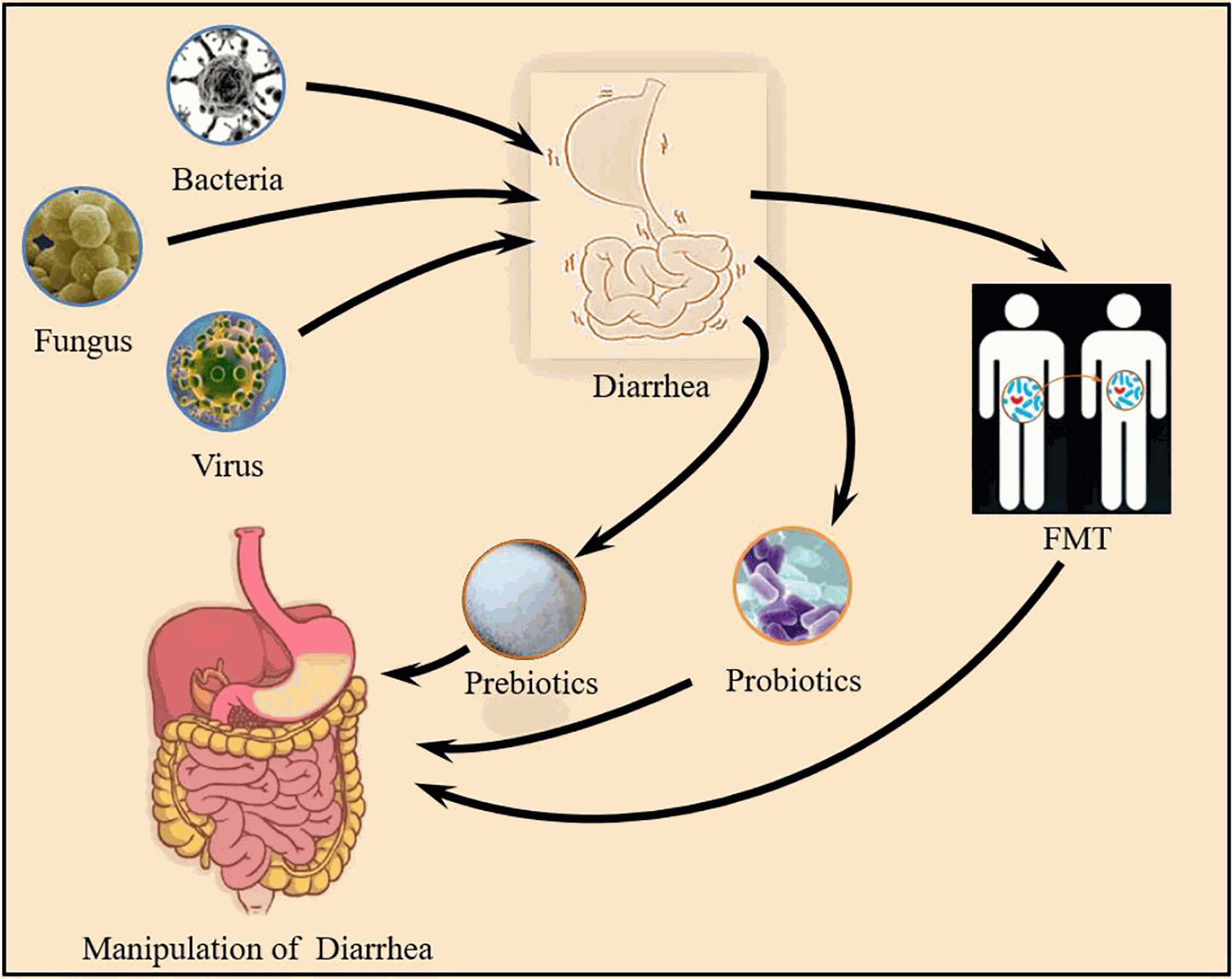
dxhh
gyb
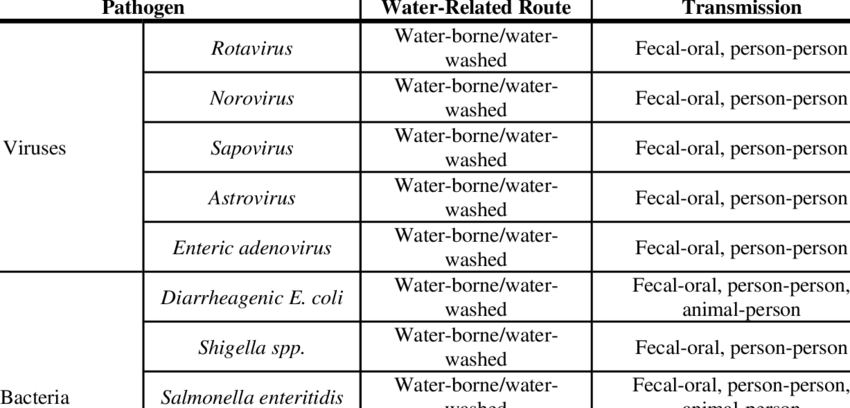
Common Pathogens causing acute diarrhea and their transmission
Specimens & Pre-analytics
- Specimens: Fresh stool or preserved stool (per collection guidelines)
- Collection: Avoid urine/water contamination; record onset date/time; ensure adequate volume
- Transport/Stability: Maintain cold chain where possible; avoid repeated freeze–thaw; aliquot for re-tests
- Pre-processing: Homogenize to minimize matrix variability; include extraction controls to detect inhibition
Workflow (Same-Day Feasible)
- Lysis & extraction (manual or automated) with an extraction control spike-in
- Multiplex amplification across 12 targets using probe-based chemistry
- Controls: Positive control (multi-analyte or panelled), non-template control (NTC), negative extraction blank
- Detection & calling: Per-target qualitative calls (Detected/Not Detected/Invalid) ± Ct values (assay-dependent)
- Review & report: Apply run-acceptance rules; issue same-day results when batching allows
Typical TAT: 2–6 hours end-to-end, platform-dependent.
Reporting & Interpretation
- Per-target results: Detected / Not Detected / Invalid, with control status
- Optional Ct metrics: Include only if validated; clarify that Ct ≠ organism viability
- Co-detections: List clearly; interpret with symptoms, duration, and exposure history to avoid over-calling colonization
- Limitations: Finite target list; stool matrix inhibition risk; no antimicrobial susceptibility
Example snippet:
- Detected: Norovirus GII (Ct 21.4)
- Not Detected: Rotavirus A, Adenovirus (enteric), Sapovirus, Astrovirus, Salmonella, Shigella/EIEC, Campylobacter, C. difficile toxin, Giardia, Cryptosporidium, E. histolytica
- Controls: Extraction IC PASS; PCR IC PASS; NTC PASS
- Interpretation: Consistent with acute viral gastroenteritis; implement contact precautions per policy; antibiotics typically not indicated for isolated norovirus.
Validation & Quality
- LoD per target in the stool matrix
- Inclusivity/Exclusivity: Circulating strains and near-neighbor cross-reactivity
- Precision: Inter-run, inter-operator, inter-instrument
- Interference: Mucus, blood, bismuth compounds, antidiarrheals
- Stability: Time/temperature studies for preserved and unpreserved stool
- EQA/PT: Enrollment and trending with CAPA for deficiencies
Clinical Value
- Rapid rule-in/rule-out for isolation and cohorting
- Stewardship: Distinguish viral from bacterial/protozoal etiologies
- Outbreak response: Same-day insights into clusters and transmission dynamics
- Reflex pathway: Negative or incongruent cases escalate to culture, antigen PCR add-ons, or mNGS